
Resources
Open data. Open minds. Explore our datasets below.
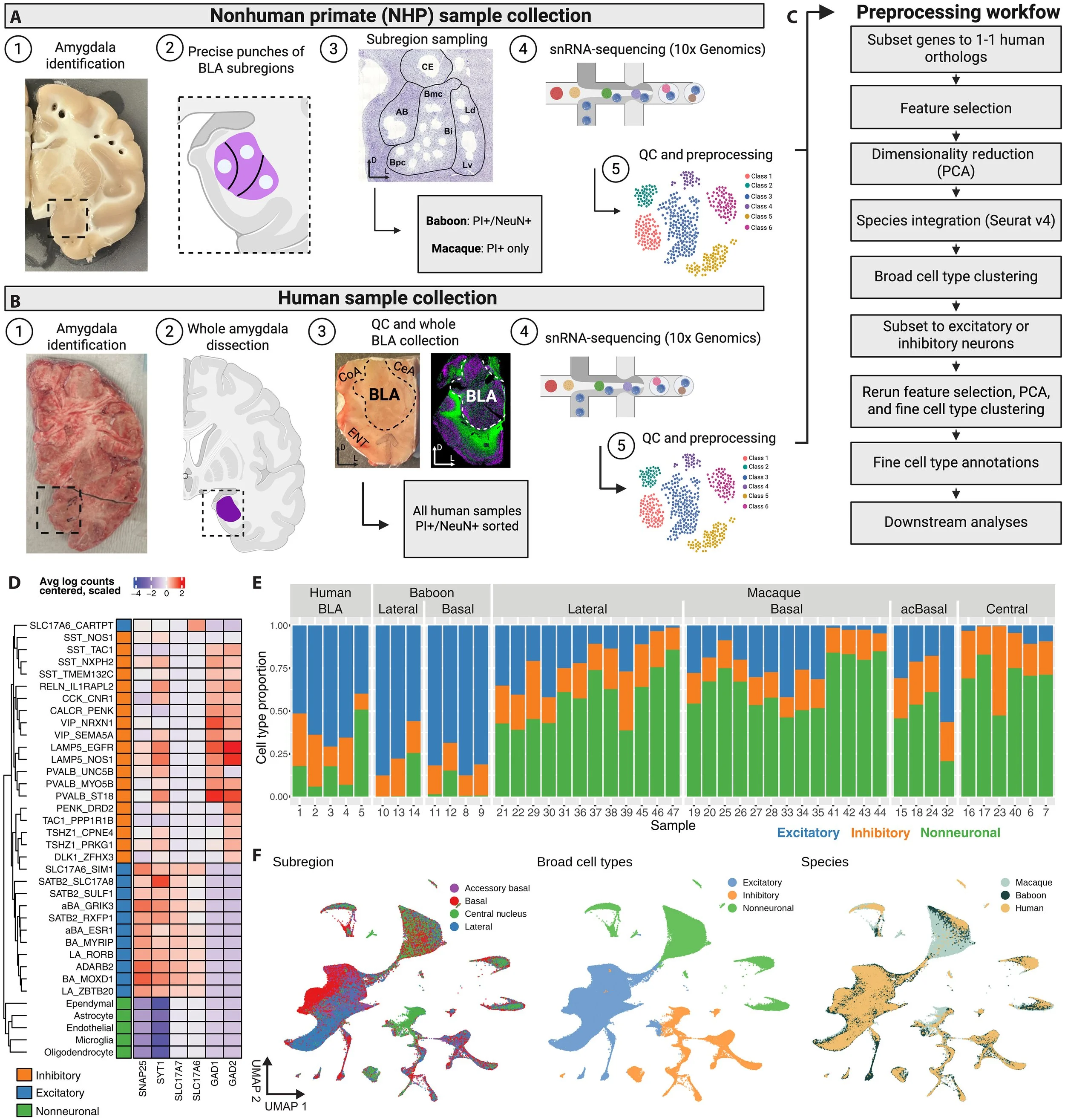
Transcriptomic diversity of amygdalar subdivisions across humans and nonhuman primates
A Comprehensive snRNA-seq Atlas of the Primate Amygdala (Human, Baboon, Macaque). Welcome to the interactive single-nucleus RNA-sequencing (snRNA-seq) atlas of the primate amygdala. This open-access resource delineates the complex neuronal organization of major amygdala subdivisions across humans, macaques, and baboons, providing a foundational tool for understanding primate brain architecture and its role in psychiatric disorders.
Key Features of the Atlas:
Cross-Species Integration: Compares cellular profiles across human and nonhuman primates (NHPs), quantifying overlaps with established rodent, macaque, and human datasets.
High-Resolution Cell Typing: Identifies five broad classes of excitatory neurons and diverse inhibitory populations (MGE, CGE, and LGE origins), highlighting substantial spatial variation across the amygdala. 32 different glutamatergic and GABAergic neuron types were identified.
Validated Marker Genes: Features robust, subdivision-specific excitatory marker genes conserved across primate species, validated via fluorescence in situ hybridization (FISH).
To promote collaboration within the broader neuroscience community, all data, code, and cloud-based exploration tools are freely accessible below.
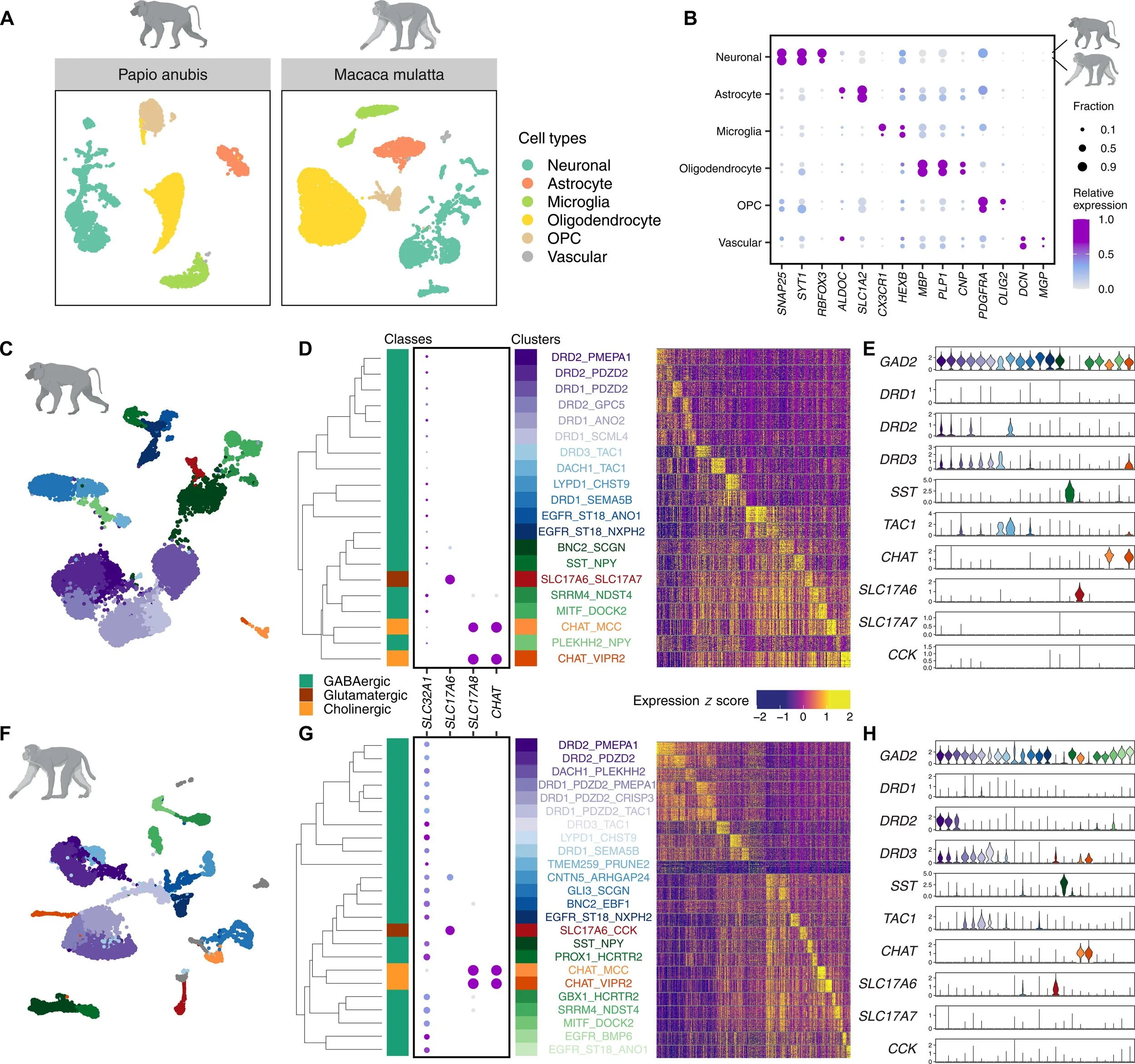
Transcriptomic landscape of mammalian ventral pallidum at single-cell resolution
A Comprehensive snRNA-seq Atlas of the Mammalian Ventral Pallidum (Macaque, Baboon, Mouse).Welcome to the single-nucleus RNA-sequencing (snRNA-seq) atlas of the mammalian ventral pallidum (VP). This open-access transcriptomic resource delineates the complex molecular taxonomy of the VP across mice, macaques, and baboons. By generating a harmonized, cross-species integration of over 87,672 neuronal nuclei, this atlas provides a consensus tool for understanding the VP, with a specific emphasis on the evolutionary specializations unique to the primate brain.
Key Features of the Atlas:
Cross-Species Integration: Successfully maps transcriptional correspondence across rodent and nonhuman primate (NHP) models. The atlas identifies 13 highly conserved neuronal clusters (comprising ~60% of VP neurons) defined by shared, functionally relevant neuropeptides and receptors (e.g., DRD2_PDZD2, CHAT_MCC).
High-Resolution Cell Typing: Moves beyond broad cell classifications to map the evolutionary divergence of the primate brain. The dataset highlights highly specialized, NHP-specific neuronal populations with no direct mouse homologs:
Enhanced Neuronal Plasticity: Identifies the NHP-exclusive DRD2_PMEPA1 cluster. Distinguished by the expression of genes critical for axonal remodeling and synaptic regulation (ACVR1, SORCS3, FOXN3), this points to increased neuronal plasticity in primates.
Novel Primate Interneurons: Characterizes the EGFR_ST18_ANO1 population, a distinct primate-specific, MGE-derived interneuron that uniquely co-expresses CGE-associated neuropeptides (TAC3, VIP).
Specialized Cholinergic Signaling:Discovers a second distinct cholinergic population unique to macaques and baboons (CHAT_VIPR2). Marked by the vasopressin receptor VIPR2, this cluster suggests a primate-specific mechanism for vasopressin-mediated excitation.
Glutamatergic Reorganization: Reveals a significant evolutionary shift in cellular composition. Glutamatergic neurons represent a notably smaller proportion of the VP in primates (2.7% in baboons, 6.6% in macaques) compared to mice (9.9%), with several rodent-specific glutamatergic clusters completely absent in NHPs.
Validated Marker Genes: Features robust, species-specific, and conserved marker genes rigorously validated via in situ hybridization. This establishes precise molecular targets for future circuit-level research and translational psychiatric investigations.
To promote collaboration within the neuroscience community, the processed gene expression metrics and cell type annotations for each individual species are freely accessible below.
Amygdala and ventral striatum make distinct contributions to reinforcement and reversal learning

This open-access dataset consists of 378,480 choices made by 11 monkeys in a probabilistic and deterministic reversal learning task. Choice and reaction time data for each choice are included.
A subset of the monkeys received excitotoxic lesions of either the amygdala or ventral striatum to determine their causal role in reinforcement learning and reversal learning.